
POJ 14(01):1-10 (2021)
ISSN:1836-3644
doi: 10.21475/POJ.14.01.21.p3141
Full Text PDF Supplementary PDF
Ghada Ahmed Abu El-Heba*
Department of Nucleic Acid and Protein Structure, Agricultural Genetic Engineering Research Institute (AGERI), Agricultural Research Centre (ARC), Giza, Egypt
Abstract
Tomato (Lycopersicon esculentum) is the main vegetal crop that has tremendous popularity around the world. Medicago truncatula NOOT gene (Mt-NOOT) encodes a BTB/POZ-ankyrin repeat protein of the NONEXPRESSOR OF PR GENES1 (NPR1 family). It was introduced into Lycopersicon esculentum (Tomato) genome. The tomato plants that ectopically expressed Mt-NOOT obtained several favorable traits and fruit quality. Heteroblasty between the transgenic and the non-transgenic tomato leaves and flower architecture were used to distinguish transgenic and wild lines. Transgenic tomato plants accumulated a significant amount of phenolic compounds and plant pigmentations compared to the wild type. On the other hand, transgenic plants acquired a considerable amount of antioxidant such as CuZnSO superoxide dismutase (SOD), tomato Catalase (CAT), and tomato Cell wall-associated peroxidase (TPX1) than the wild type. Antioxidant high content together with the high content of phenolic compounds enabled the transgenic tomato fruits to gain not only edible benefits, but also a significant higher shelf-time, extended to six months more than the wild type stored at 25°C in dark and dry condition. Surprisingly, transgenic tomato fruits did not show any rotten process during long time storage as they did not acquire any contagious microorganism. Total fruit productivity in transgenic tomato was greater than the control with an estimated ratio of 84%.
Keywords: Lycopersicon esculentum, Medicag truncatula, NOOT gene, total carotenoids, anthocyanin, phenolic compounds, macro and microelements, prolonged shelf time, high yield.
Abbreviations: CAT_Tomato Catalase, Ca+_Calcium ion, Fe++_iron ion, K+_Potassium ion,Mg++_magnesium ion, Mn++_Manganese ion, Mt-NOOT_Medicago truncatula NOOT gene, NPR1_NONEXPRESSOR OF PR GENES1 gene, Na+_Sodium ion, P+++_Phosphorus ion, SOD _superoxide dismutase, TPX1_Tomato Cell wall-associated peroxidase.
Introduction
Agricultural biotechnology is a powerful tool that can eventually contribute with the conventional method for fulfilling global need of desired quality food. Modern plant biotechnology helps us in developing genetically modified (GM) crop diversities that, in turn, can support the high demand of high yield, high nutritional balance with high tolerance to abiotic and biotic stress. The global area that accepted GM crops is expanding every year despite the concerns regarding unintended pleiotropic effects on human health. However, developing novel foods either by genetic engineering or conventional approaches is similar in the terms of potential unintended effects on human health. Tomato is one of the lowest-calorie vegetables. It has a very low-fat content and null cholesterol level. Nevertheless, it is considered as an excellent source of many vitamins, antioxidants, minerals and dietary fiber. Its antioxidant contents could be cancer-protective agents including lung, colon, breast, prostate, and pancreatic cancers. Tomato contains a considerable levels of other antioxidants like; ß-carotene, vitamin A, vitamin C, lutein, xanthine and other antioxidant enzymes (Canene-Adams et al., 2005; Uchida et al., 2010). Furthermore, tomato has a considerable level of B complex vitamins like niacin, folates, riboflavin, thiamin, as well as some essential minerals and trace elements. Increasing tomato crop productivity and enhancing its physiological characters are the main aims of this study. Several studies considered tomato plant as one of the most important economic crop, for developing transgenic tomato against various abiotic and biotic stress (Ram et al., 2019). This is achieved by understanding the plant different mechanisms and pathways that respond to such stresses like heat stress, drought stress, oxidative stress, heavy metals stress and pathogens attack (Karkute et al., 2019).
Mt-NOOT gene encodes a BTB/POZ-ankyrin repeat protein of NPR1. Systemic acquired resistance (SAR) is the Plant defense response mainly regulated by NPR1 that contains two protein–protein interaction motifs: a BTB/POZ (Broad-Complex, TraMtrack, and Bric-a-brac/Pox virus and Zinc finger) (Aravind and Koonin, 1999) and ankyrin repeats (Sedgwick and Smerdon, 1999). Upon plant’s infection, redox potential triggered by salicylic acid (SA) signal molecule accumulation allows NPR1 to enter the nucleus and interact with (TGA) transcription factors (TGACG sequence–specific binding protein), which in turn, bind to SA-responsive elements in the promoters of defense gene (PR-1 gene) (Després et al., 2003).
Mt-NOOT was isolated during mapping the symbiotic Medicago truncatula noot mutant (Couzigou et al., 2012). In the current study, Mt-NOOT was introduced into tomato genome using Agrobacterium tumefaciens to generate transgenic tomato plants aiming to assess the function of this gene in non-legume plants and test its ability to introduce some enhanced characters, which in turn, can add more edible values to tomato fruit increasing its quality and productivity. Furthermore, this study aims to prolong shelf-life time of transgenic tomato fruits without addition of preservative materials. The chemical preservatives includes nitric oxide chlorine-based solutions, hydrogen peroxide, organic acids, peroxyacetic acid, and sulphur dioxide (Kýhos et al., 2010) to the post harvested fresh crops that can add hazards to the food and affect human health. The mode of action of those chemical treatments includes ethylene bio-synthesis inhibition, respiration rate and water loss reduction, and decrease the post-harvesting diseases (Singh et al., 2013). Some other techniques were used to preserve fruits at post-harvesting such as waxing the fruit surface and storage under adjusted atmosphere (Benichou et al., 2018). However, those methods are highly expensive and toxic in some cases (Ahmed et al., 2016). Gupta et al. (2019) succeeded to improve tomato shelf- life span by manipulating polyamine content in transgenic tomato. This transgenic tomato displayed an enhancing antioxidant, ascorbic acid, and lycopene contents. However, the current study has focused on improving anti-oxidant contents in transgenic tomato as well. Previous data proved that overexpression of anti-oxidase genes including; CAT1, POD, and SOD significantly enhanced the tolerance against abiotic stresses through ROS detoxification pathway modulation (Wang et al., 2020; Muhammad et al., 2019; Ijaz et al., 2017). Other studies have manipulated CRISPR/Cas9 system for modifying one or more genes in some crops including tomato, but still the heritability of these gene alterations is remain unclear (Zhang et al., 2020).
Results
MtNOOT gene expression cassette and tomato agrobacterium tumefaciens-mediated transformation
pCambia1390::MtNOOT construct (Couzigou et al., 2012) was transformed into tomato hypocotyl explants via Agrobacterium tumefaciens. Transformation process followed by regeneration steps according to (Abu El-Heba et al., 2008).
Transgene segregation
T0 transgenic tomato plants were tested for Mt-NOOT gene integration into tomato genome. Selective PCR using noot3 and noot4 oligonucleotide primer was performed to select tomato seedlings holding Mt-NOOT. The amplified PCR fragment was 333 bp in length. Seeds from positive Mt-NOOT tomato plants were grown and allowed to self-pollination. Segregation of Mt-NOOT was confirmed in F1 via PCR giving the proper ratio of 1/1 transgenic: non-transgenic. The F2 population was grown and PCR was carried out giving 67 transgenic out of total 90 seedlings. This ratio is obeying Mendel’s segregation as shown in Figure 1A. To identify and select Mt-NOOT-Tomato lines carrying the two homozygous alleles NOOT transgene, F3 entities from each positive plant were tested separately to isolate the pure homozygous Mt-NOOT-Tomato plants.
Transgene expression and RT-PCR
Mt-NOOT transcript was confirmed by reverse transcription-PCR (RT-PCR). Extracted RNA samples from transgenic tomato leaves were analyzed by RT-PCR using noot4 and noot6 oligonucleotide primer to amplify the core region of Mt-NOOT. RT-PCR amplified a fragment of 922 bp in length, confirming the ectopic expression of Mt-NOOT in tomato tissues, while the non-transgenic tomato showed a negative result (Figure 1B).
Expression profile of SOD, CAT and TPX1 antioxidant enzymes
Tomato plant is one of the major antioxidant-containing plants and is appreciated for enormous health benefits. Antioxidant contents of tomato allows it to be one of the significant cancer prevention agent plant, decreasing harmful cholesterol levels as well. Tomato acts as an anti-aging factor and provides a worthy health condition if consumed regularly. Thus, it was so imperative to quantify antioxidant contents in transgenic tomato. Quantification of SOD, CAT and TPX1 genes expression was identified in Mt-NOOT tomato and control plants as well by Real time-PCR. Interestingly, antioxidant enzymes were up-regulated and their levels were apparently enlarged in Mt-NOOT tomato plants than wild types. SOD, CAT and TPX1 genes expression were significantly increased with a ratio of 37%, 37.5%, and 38%, respectively, in transgenic tomato plants than wild type. The average Ct for SOD in transgenic tomato was 23.62, while it was 24.73 for the control. The average Ct for CAT in transgenic tomato was 16.68 while it was 18.49 Ct for the control. The average Ct for TPX1 in transgenic tomato was 26.90 while it was 27.65 for the control. Data is shown in (Figure 2A, B and C).
Vitamin C quantification
As one of the most important antioxidants in some fruits and vegetables, Mt-NOOT transgenic tomato was subjected to Vitamin C quantification. Despite that SOD, CAT and TPX1 antioxidant enzymes were significantly higher in transgenic tomato than the control. The vitamin C quantification level was lower in transgenic tomato than the control one (Figure 3).
Mt-NOOT transgenic tomato phenotype
Transgenic tomato F3 population were grown in green house under controlled conditions in order to evaluate their phenotype and comparing it with non-transgenic tomato grown in same conditions and same developmental stage.
Shoot phenotype despite that the average fresh weights of intact transgenic plants was much higher than wild type at (p ≤0.05), the dry weights average was almost the same in both lines, assuming that transgenic tomato plants have the capability to retain abundant water contents than the non-transgenic plant (Figure 4). On the other hand, the average shoot lengths of transgenic and non-transgenic tomatoes plants were almost similar.
Leaf phenotype was one of the most characteristic phenotypes that is used to distinguish between transgenic and control phenotypes in this article, is representing the heteroblasty between the two lines. Condensation of green aerial part displayed by Mt-NOOT transgenic tomato was clearly observed in the greenhouse and the leaves number was estimated in both lines. The number of leaves originated along the primary shoot of Mt-NOOT transgenic plants was 15±2 whereas it was 11±2 in wild type. Moreover, a number of leaflets in the transgenic plant were extremely more than leaflets of the wild type. Transgenic leaflets were a deep green color, showing more wrinkles, more serrations, and tend to divide into many lobes comparing to the wild type at the same developmental stage. In contrast, wild type plants appeared smooth leaf with smooth margins and the leaflet was intact without lobes. Some obvious complexity was observed in transgenic tomato leaves, whereas some terminal leaflet was fused with lateral one. In some others the terminal leaflet was missing. Heteroblasty between Mt-NOOT transgenic line and the non-transgenic line is shown in (Figure 5 and Supplementary Figure 1).
Flower and fruit phenotype although flowers of transgenic tomato are radially symmetrical and the number of sepals, petals, and Stamen are similar to wild type, transgenic tomato flowers are showing a pointed needle shape petals as a diverse phenotype (Supplementary Figure 2 A). Because flowering is the major stage of fructification, flower number and flower phenotype were important topics in phenotype description. Furthermore, the ratio of fruit formation to the first blooms was essential in demonstrating the transgenic yield of tomato plants. Transgenic tomato plants displayed a significantly higher number of flowers with about 15±2 flowers against 4±2 flowers in wild type. Furthermore, the ratio of fruits formed from first flowers was 100% per each bloom in the case of transgenic plants, while it was 50% in control (Supplementary figure 2B & C). So the productivity of the total fruit in transgenic tomato was greater than the control with an estimated ratio of 84%. Fruit dimension and outermost smooth texture of Mt-NOOT-Tomato were similar to those of non-transgenic tomato and there is no dissimilarity displayed by both lines except the needle shape like sepals of the transgenic flower (Supplementary Figure 2D & E).
Tomato fruit ripening and storage
Many events can affect fruitlet progress and consequently fruit size and quality. Breaker stage is when up to 10% of the tomato’s surface begin shifting their green color to red or pink color at the blossom end. Harvesting tomato fruits at five days-post breaker stage was to ensure that transgenic and non-transgenic tomatoes are fully mature and their flavor and pigmentations are completely developed, meaning the ethylene gas is produced internally to enhance ripening process. Two weeks post harvested spoiling process and the rotten feature was observed on the non-transgenic tomato stored at 25°C dark dry condition as described in materials and methods section. The transgenic tomato fruits displayed a healthy feature without contamination or wilting appearance stored in the same condition. Interestingly, shelf life duration of transgenic tomato was 6±1 months comparing to only 2±1 weeks in non-transgenic tomato fruit without adding any food preservation substances (Supplementary Figure 2 F, G, H).
Physiological characterization
Determination of Mt-NOOT-Tomato plant pigmentation
Chlorophylls, total carotenoids, and anthocyanin are three kinds of pigments existing in plants and their production quantity determines leaves and fruits color. Due to the observed dark green color leaves of Mt-NOOT-Tomato, total chlorophyll, total carotenoids, and anthocyanin pigments were estimated in transgenic and non-transgenic tomato plants as well. Chlorophyll A did not show any significant difference between their content in transgenic and non-transgenic at (p=0.2) while chlorophyll B was significantly higher than the control at (p≤ 0.05). On the other hand, total carotenoids and anthocyanin were significantly higher in transgenic tomato than the control at (p ≤ 0.05). Chlorophyll, carotenoids, and anthocyanin contents estimation is represented in Figure 6A. The high contents of these pigments comparing to the wild type can explain the darker color observed in Mt-NOOT-Tomato.
Quantification of phenolic compounds in Mt-NOOT-Tomato
Phenolic compounds are one of the largest class of plant secondary metabolites. Quality and flavors property of fruits and vegetable depend on phenolic compounds contents and they are responsible for red fruit’s color as well. Several health benefits can be achieved through fruits and vegetables consumption due to phenolic compounds and their derivatives. Quantification of phenolic compounds in Mt-NOOT-Tomato is very essential to determine whether the dark green color of the leaves is linked to the enlarged quantity of such compounds or not. Spectrophotometer was used to estimate the total contents of the phenolic compound calorimetrically. Significant difference was observed between Mt-NOOT-Tomato and control tomato as they accumulate a large amount of phenolic compounds than Control at p=0.00 as shown in (Figure 6B).
Quantification macro-elements and micro-elements in Mt-NOOT-Tomato
Plant needs essential nutrients to survive and blossom. Essential nutrients includes macronutrients that plant needs in great quantity and micronutrients that plant needs in tiny amounts. These nutrients beside the sunlight and water are crucial for a plant's survival. Some essential macronutrients and micronutrients contents were subjected to measurements in Mt-NOOT-Tomato plants as well as control plants. K+ and P+++ macroelements concentrations were significantly higher than wild type tomato at (p ≤ 0.004), and (p ≤0.001), respectively while Ca+, Mg++ concentrations were significantly lower than wild type tomato at (p=0.00), and (p≤0.001) respectively. Fe+++, Mn++ were significantly higher than wild type tomato at (p=0.00), and (p ≤0.001), respectively, while Cu++ concentration was significantly lower than wild type tomato at (p=0.00), and (p≤0.05) as shown in (Figure 7).
concentration was significantly lower than wild type tomato at (p=0.00), and (p≤0.05) as shown in (Figure 7).
Discussion
The significance of ideal nutrition for human wellbeing and development is much perceived. Therefore, the major objective of scientists is to discover a way to sustain high yield even under harsh conditions and engineering crops with improving nutritional quality as well. Genetic modified (GM) crops are a powerful tool than conventional approaches for satisfying the worldwide need of high nutrition value foods. GM crops can provide biotic and abiotic stress tolerance as well. In the last years, the availability of thousands of genes has enabled us to improve both nutrition value, yield and to obtain more tolerant verities for the most economic importance crops factor (Datta 2012; KaMthan et al., 2012). Çolak (2020) determined that the crop nutritional value depends not only on proteins, lipids and sugar contents but also on compounds such as antioxidants, lycopene, vitamin C, and β-carotene. So tomato as the most popular consumed fruit worldwide has been considered in this study. Tomato can be edible as raw food, cooked food, or even processed in enormous industrial products. On the other hand, improving fruit quality, productivity, increasing shelf- time, and enhancing its minerals and antioxidants contents is such great demand.
Incredibly, tomato ectopically expressing Mt-NOOT gene thrived to provide healthy tomato plants with more nutrients contents such as minerals, antioxidant carotenoids, and phenolic contents. Moreover, the developed transgenic tomato fruits achieved extremely long shelf life and protection against various pathogens despite that wild type tomato is metabolically very active and has a short shelf life. Mt-NOOT gene shows 77% identity and 85% similarity to NPR-like BLADE-ON-PETIOLE1/BLADE-ON-PETIOLE2 (BOP2) (Ha et al., 2004; Norberg et al., 2005) and is the closest homologs to Arabidopsis BOP1/BOP2 genes. Previously, it was suggested that Bop and NPR1 genes have distinctive biological functions (Couzigou et al., 2012) depending on the fact that bop1 bop2 mutant did not alter Arabidopsis response to pathogens but rather it altered leaves and flower phenotype (Hepworth et al., 2005), whereas NPR1 overexpression enriches plant defense in Arabidopsis against the different pathogens (Cao et al., 1998; Friedrich et al., 2001). NPR1 contributes in plant defense mechanism via salicylic acid signals (Pieterse and Van Loon, 2004). In here, transgenic tomato’s phenotype was precisely analyzed to evaluate the Mt-NOOT gene function in non-legumes plant and to answer the question if NOOT gene is involving in defense mechanism in non-legume and if it can actively influence plant phenotype.
The most remarkable observation in transgenic tomato plants was leaf phenotype. The Mt-NOOT tomato showed heteroblasty when compared to the control one. KNOTTED1 LIKE HOMEOBOX (KNOX1) gene expression is maintained in shoot apical meristem (SAM) affecting its formation. KNOX1 is commonly down-regulated through the switching state from the meristem stage to the primordial stage. In simple leaf, this down-regulation is permanently stood like in Arabidopsis, while in tomato as it has a compound leaf, it is re-developed in the primordial establishment. Overexpression of KNOX1 shows plants with wrinkle and lobed leaves (Uchida et al., 2010). This article showed that the transgenic tomato showed leaf architecture complexity, wrinkled leaflet, serrated margins, and more deep lobes while wild-type was smooth margins, less in both wrinkled and lobes with simple architecture. We assumed that Mt-NOOT overexpression is driving KNOX1 up-regulation causing such phenotype. During leaf development, the primordia are adopted distinct progressive patterning through central-lateral, abaxial-adaxial, and proximal-distal axes. The way of cell division and expansion defines both leaf size and shape (Hepworth et al., 2005). Change in the leaf proximal-distal axis was detected in Mt-NOOT-Tomato as it was longer and wider than wild type leaf. Thus, results assumed that Mt-NOOT is orthologous to ASYMMETRIC LEAVES1 (AS1), AS2 and BOP1 which encodes a MYB transcription factor (Byrne et al., 2000; Ori et al., 2000; Semiarti et al., 2001; Sun et al., 2002), a LOB domain/Leu zipper transcription factor (Iwakawa et al., 2002; Lin et al., 2003), and NPR1-like transcription factor (Ha et al., 2003, 2004) respectively. They are controlling proximal-distal modeling.
The plant’s pigmentation, chlorophylls, total carotenoids, and anthocyanin were quantified in transgenic tomato. Chlorophyll B, total carotenoids and anthocyanin were found to be enormously greater in transgenic tomato than the wild type tomato added more quality for the Mt-NOOT-Tomato plants. Chlorophyll A and B are the photosynthesis pigments, produced in leaf chloroplasts. Chlorophylls are eventually broken down at the end of leaf life and the nitrogen is absorbed by the plant. Total carotenoids are kind of antioxidant pigments, synthesized in plant plastids. Total carotenoids act as vision protecting agent fighting cellular damage (Armstrong et al., 1996). Carotenoids support the cardiovascular system, liver, breast, colon, lungs, and prostate health. The anthocyanin offers anti-viral, anti-inflammatory, and anti-cancer agents and provides antioxidant benefits to human health. On the other, Phenolic compounds displayed a high contents in transgenic tomato.
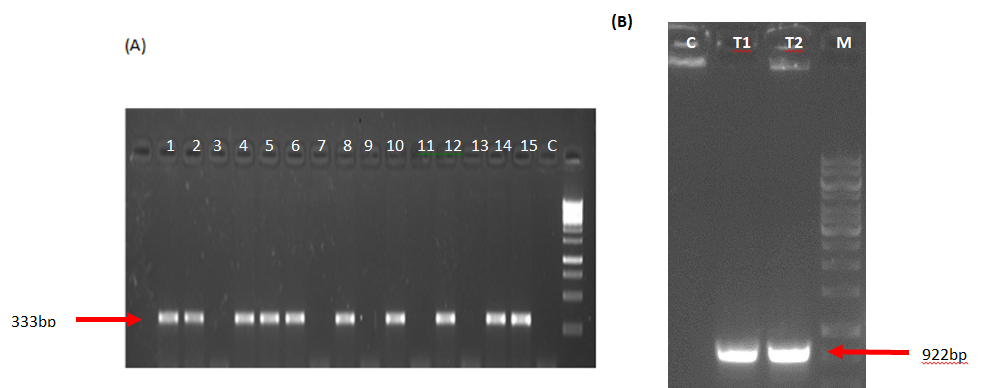
Figure1. Mt-NOOT Segregation and determination of its transcript (A) Segregation of Mt-NOOT gene in F2 transgenic tomato giving the proper percentage of 3:1 transgenic: non-transgenic ratio obeying Mendel’s low of segregation. About 15 individuals from F2 transgenic Tomato were chosen randomly for the experiment, noot3 & noot4 oligonucleotide pair was used to amplify a NOOT gene band of 333bp including part of exon1 while (B) the oligonucleotide prime pair noot4 & noot6 was used to amplify the core gene and confirm transgene expression of Mt-NOOT inside Tomato tissue giving rise to a fragment of 922bp in length. Control healthy Tomato was used as a control in the both amplification experiments.
A protecting role of phenolic compounds against carcinogenic agents is maintained by many researches (Basli et al, 2017).
Essential minerals like potassium, phosphorous, iron and manganese were significantly increased in transgenic plants. The main role of potassium is regulation of osmotic pressure in plants. Plants suffering from potassium deficiency are unable to tolerate water stress. This phenomenon can explain the higher amount of water contents in Mt-NOOT-Tomato than the wild type. Since the transgenic tomato obtains more potassium content, water retention is improved. Phosphorus may enhance the fruit quality and increase plant disease resistance as well. Elevated iron contents in Mt-NOOT transgenic tomato improves other nutrient absorption and helps to carry out all enzymatic reactions in plants such as peroxidase, nitrogenase, catalase, aconitase, etc. Chlorophyll contents was significantly higher in Mt-NOOT transgenic tomato accompanied to the expanding contents of manganese microelement, as manganese concentration is directly proportional to chlorophyll concentration. An ideal manganese concentration may counteract with bad consequence of low quality aeration.
High tomato fruit yield depends mainly on high blossoms production by the healthy plant containing adequate minerals and nutrients supplement. Transgenic tomato plants showed significant high levels of pigments, phenols, antioxidants contents and some macro and microelements. Their flower number was considerably more than the non-transgenic tomato. This is assumingly due to the high phosphorus contents of the Mt-NOOT-Tomato. The phosphorus is involved in stimulating bud development and blooming process. This fact can explain the condensation of the vegetative part in transgenic plant than the control. Thi Man Le (2020) showed that magnesium supplement to tomato plants can cause a reduction in plant height but increased fruit yield. Fortunately, almost 100% flowers produced by Mt-NOOT-Tomato were pollinated and fruits set compared to only about 50% in control plants. Altogether the higher flower production and the higher fruit productions of transgenic tomato enabled higher fruit yield (84%), than the wild type tomato.
Fresh crops post-harvest treatments are used to minimize the losses in the nutritional and value attributes as it slows down physiological processes in crops such as senescence, respiration, and ripening (Lopez-Galvez et al., 2013). However, using manufacturing materials for such purpose is not appreciated for their biohazard effects. Thus, it was essential to find a natural approach to increase the shelf lifetime of fresh fruits and vegetables. Hence, a Shelf life is such a significant value trait for various fruit including tomato. In this study, fruits of transgenic tomato ectopically expressing Mt-NOOT gene were proven to reduce the prevalence of various pathogens attack and have an extraordinarily increasing shelf-life duration. This indicates that NOOT gene has a perspective role in extremely extending shelf- life of tomato fruits for an extended period of 120 days more than control tomato. This long shelf time duration of transgenic tomato proposedly is due to significantly high levels of antioxidants contents such as, cytosolic CuZnSO superoxide dismutase, catalase, and cell wall-associated peroxidase. Bhagwan et al. (2000) reported that post-harvest treatment with antioxidants such as benzyladenine a sodium benzoate, and ascorbic acid improve shelf life and several phytochemical possess. The other suggestion is that the prolonged shelf time of transgenic tomato is due to the increasing level of plant pigmentation like total carotenoids and anthocyanin. This suggestion is agreed with Zhang et al. (2013) who reported that natural pigments enrichment particularly anthocyanin in tomato can prolong shelf life significantly. They proved that tomato ripening are repressed and its susceptibility to the most vital postharvest pathogen, Botrytis cinerea, is reduced by the accumulation of anthocyanin pigment. Amazingly, after the six-month shelf time duration, transgenic tomato fruits were standing healthy without any contamination due to the high contents of phenolic compounds as well as total carotenoids and anthocyanin in Mt-NOOT transgenic tomato.
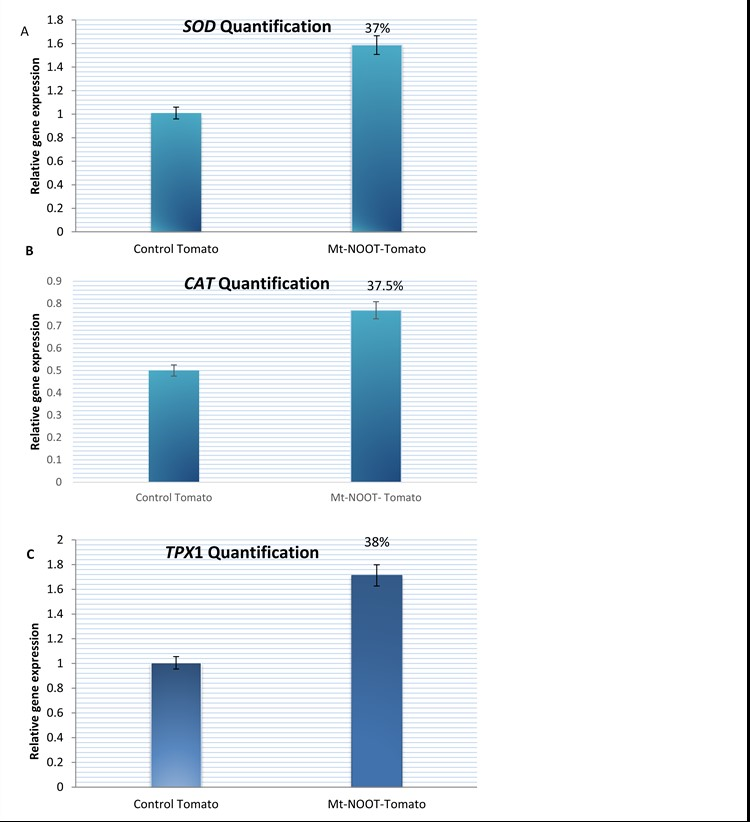
Figure 2. Quantification real time PCR for (A) SOD, (B) CAT and(C) TPX1 genes was performed in Mt-NOOT-Tomato plants and control plants as well using three sets of specific oligonucleotide primers. Actin gene was used as the endogenous stably expressed gene. SOD, CAT and TPX1 antioxidant genes’ expressions are increased with a ratio of 37%, 37.5%, and 38% than the control.
Materials and Methods
Plant source
In this study, tomato (Lycopersicon esculentum) cv. CastleRock seeds were obtained commercially from Agricultural Research Center and used as a plant material. The MS medium was supplemented with 3% sucrose and 0.25% phytagel with a pH adjusted to 5.8 before autoclaving was used as growth media. In vitro growth chamber with a growth condition of 25°C and 8/16 hr (dark/light) photoperiod was used for incubating all regeneration culture media.
Plant expression vector and agrobacterium transformation
A fragment of 2811 bp NOOT was cloned in modified pCAMBIA1390 vector under the control of cauliflower mosaic virus 35S promoter (Couzigou et al., 2012). The construct was transformed into Agrobacterium strain GV3101 (Koncz et al., 1989) via electroporation (Sukharev et al., 1992) at 400 V, 25 mF and 2.5 kV. The transformed cells were selected on LB agar medium supplemented with 50 mg/ml streptomycin and 50 mg/ml kanamycin. Tomato hypocotyl was used as explant source for transformation followed by the regeneration process as described by (Abu El-Heba et al., 2008).
Genetic inheritance and DNA extraction
To select transgenic tomato entities holding the two homologues allele of Mt-NOOT, F0, F1, F2, and F3 populations were grown in green house. Leaves from one month old seedlings were subjected to DNA extraction using QIAGEN Genomic DNA isolation kit according to manufacturer’s instruction. The DNAs from all transformed seedlings were used for PCR reaction using EmeraldAmp PCR Master MIX, TAKARA. Oligonucleotide primers pair noot3: 5’- GCGGCTGCAAGAGTATCAAGAGC-3’ and noot4: 5’- CAGCGTCGAAGGACGTTTAGTCC-3’. The denature temperature was 95°C for 5 min, annealing 60°C for 1 min and extension was at 72°C for 30 sec followed by the final extension at 72°C for 7 min. Products were visualized on 1% agarose gel using a 1 kb DNA ladder. Tomato entities with positive PCR result were grown in controlled green house and allowed for self-pollination. Seeds were harvested and grown at same growth conditions. F2 seedlings were subjected to DNA extraction and PCR amplification to evaluate the segregation pattern of NOOT allele. Entities homozygous for the NOOT allele were grown and allowed to self-fertilize to produce F3 population.
Detection of Mt-NOOT expression via RT-PCR
Total RNA was extracted from F3 entities of Mt-NOOT-tomato transgenic population as well as control tomato plant (five replicates for each). About 100 mg leaves was used using Qiagen Kit (RNeasy Mini Kit) according to the manufacturer’s instructions. DNase1 RNase free kit (Thermo Scientific) was used to remove all DNA contents. Quantity of the recovered RNAs and their purity were measured via spectrophotometry absorbance at 260 and 280 nm. All steps were achieved under sterile conditions to reduce contamination risks. One step RT-PCR was performed according to manufacturer’s instruction using SuperScript-III with the Platinum Taq DNA Polymerase. Oligonucleotide pair; noot4 and noot6: 5’-GGGTCAGCGTGGTGGTCGAGCAA-3’ was used for NOOT expression detection using PCR conditions as described above.
Quantitative Real time-PCR for SOD, CAT, TPX1
Polyadenylated mRNA was extracted from F3 entities Mt-NOOT-Tomato five samples (5 biological replicates) each tested triple times using Qiagen Kit (Oligotex mRNA Mini Kit), and reverse transcribed into cDNA.
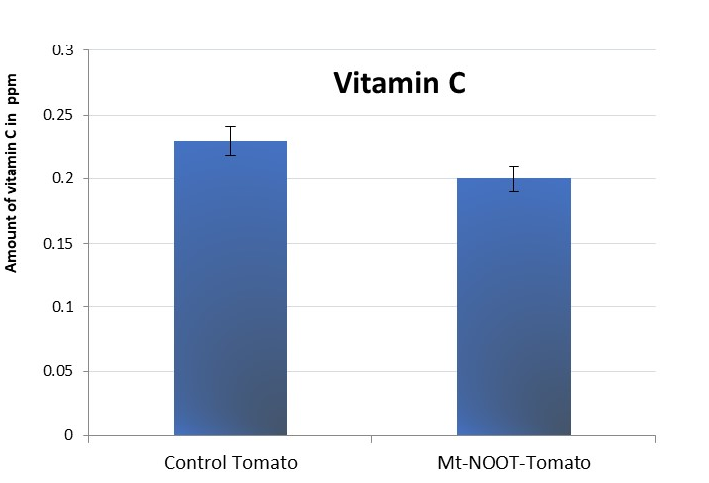
Figure 3. Vitamin C quantification in transgenic and non-transgenic Tomato showed an increased level in control plants than the Mt-NOOT transgenic Tomato (30 biological replica were used for each line).
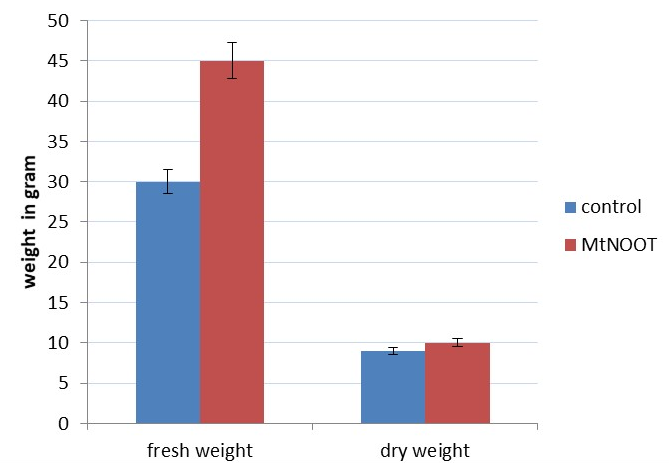
Figure 4. Differences in fresh and dry weight between Mt-NOOT transgenic Tomato and control Tomato demonstrating the high capability of water retaining by transgenic Tomato comparing to the wild type (30 biological replica were used for each line). Fresh weight of transgenic Tomato was significantly higher than the wild type at (p ≤0.05).
The reaction of 20 μL contained polyadenylated mRNA, 250 ng random hexamer primer, dNTPs 0.5mmol/L, 1× First Strand Buffer, DTT 10 mmol/L, 30 units RNase inhibitor, SuperScript-III reverse transcriptase, and DNase RNase Free ultra-pure distilled water (Invitrogen, Life Technologies, Inc.). Samples were incubated at 42°C for 50 min and reactions were inactivated at 70°C for 15 min. Real time-PCR was accomplished via AB7300 Real-Time PCR system (AppliedBiosystems, USA). Specific oligonucleotide primers SOD, CAT, and TPX1 (El-Gaied et al., 2013) for tomato cytosolic CuZnSO superoxide dismutase, tomato Catalase, and tomato Cell wall-associated peroxidase were respectively used for the antioxidant quantification of Mt-NOOT transgenic tomato plants with expected products of 153 bp, 145 bp, and 112 bp, respectively. Actin gene (ID: 101260631 NCBI), the endogenous stably expressed gene, was amplified as control for quantity and quality of input RNAs. The positive control for this assay was the RNA from healthy non-transgenic tomato plant. Reactions contain samples cDNA, oligonucleotide primers, Invitrogen Platinum Taq DNA Polymerase, SYBR Green qPCR Master Mix Thermo Scientific, ROX dye as a reference dye and the volume was divided into 3 tubes each for 50 μL. 96-Well reaction plate was sealed with Optical Caps (MicroAmp, Applied Biosystems) and placed in AB7300 Real-Time PCR system (Applied Biosystems). PCR program initiated according to manufacturer’s instructions. Three technical replicates were performed for each sample.
Morphological characterization
Tomato plants ectopically expressing Mt-NOOT gene as well as control tomato plants were grown in green house at 25°C in 16/8 hr (light/dark) photoperiod and the phenotype were detected. 30 biological replicates were tested for each line. Shoot lengths, fresh and dry weights of flowers and fruit number were observed.
Fruit storage condition
Fruit shelf life time duration was verified in transgenic and non-transgenic tomato. Tomato fruits were harvested five days post-breaker and the harvesting process was continued until the plants stop giving fruits. Watering tomato plants was halted three days pre-harvesting to enhance the ripening process. The collected fruits were completely dry surfaced and stored at 25°C dark dry room. Tomato fruits were arranged flipped upside down in vented plastic harvested bins.
Physiological characterization
Determination of plant pigments in Mt-NOOT-Tomato
About 100 mg of fresh leaves from transformed line and control line, each with 30 biological replica were grounded using liquid nitrogen. Total chlorophyll, carotenoids and anthocyanin pigments were extracted by 80% acetone containing 1mM potassium hydroxide, followed by centrifugation at 10,000 rpm for 10 min. spectrophotometer (Ependorf) device was used for supernatant quantification (Schelbert et al., 2009). Chlorophyll, total carotenoids and anthocyanin amounts were estimated according to (Lichtenthaler and Buschmann 2001).
Phenolic compounds Quantification in Mt-NOOT-Tomato
Total phenol content was estimated via Folin Ciocalteu’s protocol according to (Patel et al., 2010). 30 biological replications were used for each line, 1 g of each sample was extracted in 3ml of 95% ethanol and kept in dark at 4°C/24hrs, and then samples were centrifuged for 5 min at 13000 rpm. 1 ml supernatant was mixed with 5 ml distilled water, 0.5 ml Folin Ciocalteu reagent, 1.5 Na2CO3 20% and the volume was adjusted to 10 ml and incubated for 2 hrs at room temp. Spectrophotometer (Ependorf) was used to detect samples reading at 765nm. Total phenolic contents was calculated using standard curve generated with 10 to 100 mg/ml concentrations of salicylic acid.
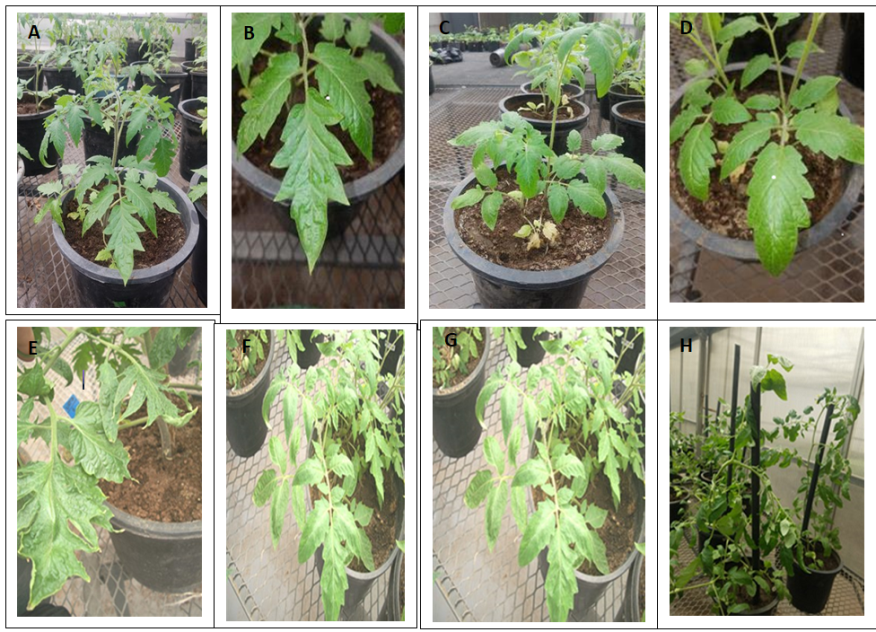

Figure 5. Tomato plants have compound leaves made up of leaflets that are scattered along leaf rachis, some of these leaflets are compound. The entire leaf is attached to stem by petiole while leaflets are connected to rachis by petiole. Heteroblasty was shown between Mt-NOOT transgenic Tomato leaves and control one. Transgenic Tomato leaflets are deep in green color, wrinkled, serrated and divided into many lobes (A) & (B) & (E). Control Tomato plants showing smooth leaf texture, less serration and less lobes (C) & (D) & (F). Overall Mt-NOOT- transgenic Tomato plants are showing high dense vegetative Ariel part due to higher number of leaflets per each leaf (G) comparing to the wild type plants (H).
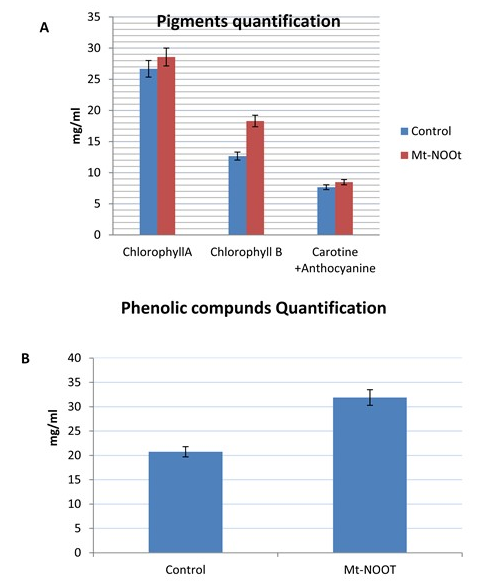
Figure 6. Plants pigmentation and phenolic compound quantification were estimated in transgenic Tomato ectopically expressed Mt-NOOT gene (A), Chlorophyll A don’t show any significant difference between transgenic and non-transgenic at (p≤0.2) while chlorophyll B was significantly higher in transgenic than the control at (p≤ 0.05), Carotenoids and anthocyanin were significantly higher in transgenic than the control at (p ≤ 0.05). In a similar manner, Transgenic Tomato is accumulate large amount of phenolic compounds than Control at p≤0.00 (B).
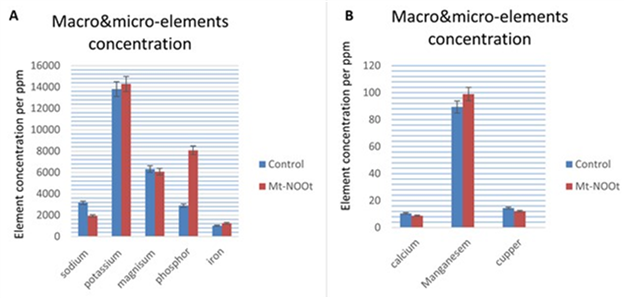
Figure 7. Plants pigmentation and phenolic compound quantification were estimated in transgenic Tomato ectopically expressed Mt-NOOT gene (A), Chlorophyll A don’t show any significant difference between transgenic and non-transgenic at (p=0.2) while chlorophyll B was significantly higher in transgenic than the control at K^+, P^(+++) macro-elements concentrations were significantly higher than wild type Tomato at (p ≤ 0.004), and (p ≤0.001), respectively while Ca^+,Mg^(++) concentrations were significantly lower than wild type Tomato at (p≤0.00), and (p≤0.001) respectively. Fe^(+++) [Mn] ^(++) were significantly higher than wild type Tomato at (p=0.00), and (p ≤0.001), respectively while [Cu] ^(++) concentration was significantly lower than wild type Tomato at (p≤0.00), and (p≤0.05). (p≤ 0.05), Carotenoids and anthocyanin were significantly higher in transgenic than the control at (p ≤ 0.05). In a similar manner, Transgenic Tomato is accumulate large amount of phenolic compounds than Control at p≤0.00 (B).
Ion quantification in Mt-NOOT-Tomato
Ten replica of 15gm dry weight of both Mt-NOOT-Tomato and control plants were subjected to K+, Mg++, Na+, Ca+, P+++, Mn++ and Fe++ ions concentration measurement. The measurements were carried out in Regional Center for Food and Feed, ARC, Giza, Egypt.
Statistical analysis
Mean value of 30 replica per each line were calculated and followed by analysis using SSPS- One-Way ANOVA method.
Conclusion
Tomato plants ectopically expressing Mt-NOOT obtained several favorable traits and fruit quality and quantity. Anti-oxidant agents like; tomato cytosolic CuZnSO superoxide dismutase, tomato Catalase, and tomato Cell wall-associated peroxidase were higher in transgenic with a ratio of 37%, 37.5%, and 38%, respectively, than the wild type. In a similar manner, the transgenic tomato plants accumulated a significant amount of phenolic compounds, and plant pigmentation including; chlorophyll, total carotenoids, and anthocyanin comparing to wild type. In contrast, the transgenic tomato showed a declining level of vitamin C than the control tomato. Essential elements like K+, P+++, Fe+++, Mn++ were significantly higher in transgenic tomato than wild type tomato. Interestingly, shelf life duration of transgenic tomato was 6±1 months, compared to only 2±1 weeks in non-transgenic tomato fruit without adding any food preservation substances.
Acknowledgments
The author appreciates Ms. Asmaa Hamdy for her assistance in dealing with transgenic tomato and Dr. Mohamed Abd El-Satar for performing Real time PCR.
References
Abu El-Heba GA, Hussein GM, Abdalla NA (2008) A rapid and efficient tomato regeneration and transformation system. Agric For Res. 58:103–110.
Ahmed MF, Islam MZ, Sarker MSH, Hasan SMK, Mizan R (2016) Development and performance study of controlled atmosphere for fresh tomato. World J Eng Tchnol. 4:168–175.
Aravind L, and Koonin EV (1999) Fold prediction and evolutionary analysis of the POZ domain: Structural and evolutionary relationship with the potassium channel tetramerization domain. J Mol Biol. 285:1353–1361.
Armstrong GA, Hearst JE. (1996) Carotenoids 2: Genetics and molecular biology of carotenoid pigment biosynthesis. FASEB J. 10:228-37.
Basli A, Belkacem N, Amrani I (2017) Health benefits of phenolic compounds against cancers. In: Phenolic compounds. Soto-Hernández M, Palma-Tenango M, Garcia-Mateos MR (eds) IntechOpen. London.
Benichou M, Ayour J, Sagar M, Alahyane A, Elateri I, Aitoubahou A (2018) Postharvest technologies for shelf life enhancement of temperate fruits. In: Mir S, Shah M, Mir M (eds) Postharvest biology and technology of temperate fruits. Springer, Cham.
Bhagwan A, Reddy YN, Rao PV, Mohankumar KC (2000) Shelf life extension of tomato fruits by postharvest antioxidant application. J Appl Hortic. 2: 88-91.
Byrne ME, Barley R, Curtis M, Arroyo JM, Dunham M, Hudson A, Martienssen RA (2000). Asymmetric leaves1 mediates leaf patterning and stem cell function in Arabidopsis. Nature. 408: 967–971.
Canene-Adams K, Campbell JK, Zaripheh S, Jeffery EH, Erdman JW, Jr (2005) The tomato as a functional food. J Nutr. 135:1226–1230.
Cao H, Glazebrook J, Clarke JD, Volko S, Dong X (1998) The Arabidopsis NPR1 gene that controls systemic acquired resistance encodes a novel protein containing ankyrin repeats. Cell. 88:57–63.
Çolak NG, Eken NT, Ülger M, Frary A, Doğanlar S (2020) Mapping of quantitative trait loci for antioxidant molecules in tomato fruit: Carotenoids, vitamins C and E, glutathione and phenolic acids. Plant Sci. 292:110393.
Couzigou JM, Zhuko V, Mondy S, Abu el Heba G, Cosson V, Noel Ellis TH, Ambrose M, Wen J, Tadege M, Tikhonovich I, Mysore KS, Putterill J, Hofer J, Borisov AY, Rateta P (2012) NODULE ROOT and COCHLEATA maintain nodule development and are legume orthologs of Arabidopsis BLADE-ON-PETIOLE Genes. Plant Cell. 24:4498–4510.
Datta A (2012) GM crops: dream to bring science to society. Agric Res. 1: 95-99.
Després C, Chubak C, Rochon A, Clark R, Bethune T, Desveaux D, Fobert PR (2003) The Arabidopsis NPR1 disease resistance protein is a novel cofactor that confers redox regulation of DNA binding activity to the basic domain/leucine zipper transcription factor TGA1. Plant Cell. 15:2181-2191.
Friedrich L, Lawton K, Dietrich R, Willits M, Cade R, Ryals J (2001) NIM1 overexpression in Arabidopsis potentiates plant disease resistance and results in enhanced effectiveness of fungicides. Mol Plant Microbe In. 1:1114–1124.
Gupta A, Pandey R, Sinha R, Chowdhary A, Pal RK, Rajam MV (2019) Improvement of post-harvest fruit characteristics in tomato by fruit-specifc over-expression of oat arginine decarboxylase gene. Plant Grow Regul. 88:61–71.
Ha CM, Kim GT, Kim BC, Jun JH, Soh MS, Ueno Y, Mahida Y, Tsukaya H, Nam HG (2003) The BLADEON-PETIOLE1 gene controls leaf pattern formation through the modulation of meristematic activity in Arabidopsis. Development. 130:161–172.
Ha CM, Jun JH, Nam HG, Fletcher JC (2004) BLADEON-PETIOLE1 encodes a BTB/POZ domain protein required for leaf morphogenesis in Arabidopsis thaliana. Plant Cell Physiol. 45:1361–1370.
Hepworth SR, Zhang Y, McKim S, Li X, Haughn GW (2005) BLADE-ON-PETIOLE-dependent signaling controls leaf and floral patterning in Arabidopsis. Plant Cell. 17:1434–1448.
Ijaz R, Ejaz J, Gao S, Liu T, Imtiaz M, Ye Z, Wang T (2017) Overexpression of annexin gene AnnSp2, enhances drought and salt tolerance through modulation of ABA synthesis and scavenging ROS in tomato. Sci Rep. 7:12087.
Iwakawa H, Ueno Y, Semiarti E, Onouchi H, Kojima S, Tsukaya H, Hasebe M, Soma T, Ikezaki M, Machida C, Machida Y (2002) The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana, required for formation of a symmetric flat leaf lamina, encodes a member of a novel family of proteins characterized by cysteine repeats and a leucine zipper. Plant Cell Physiol. 43:467–478.
KaMthan A, KaMthan M, Azam M, Chakraborty N, Chakraborty S, Datta A (2012) Expression of a fungal sterol desaturase improves tomato drought tolerance, pathogen resistance and nutritional quality. Sci Rep. 2: 951
Karkute SG, Krishna R, Ansari WA, Singh B, Singh PM, Singh M, Singh AK (2019) Heterologous expression of the AtDREB1A gene in tomato confers tolerance to chilling stress. Biol Plantarum. 63: 268-277
Koncz C, Martini N, Mayerhofer R, Koncz-Kalman Z, K€orber H, Redei GP, Schell J (1989) High-frequency TDNA-mediated gene tagging in plants. Proc Natl Acad Sci USA. 86: 8467–8471.
Kýhos K, Průchová J, Novotná P (2010) Decontamination of cut carrot by persteril® agent based on the action of peroxyacetic acid. Czech J Food Sci. 28: 564-571.
Lichtenthaler HK, Buschmann C (2001) Extraction of photosynthetic tissues: chlorophylls and carotenoids. Curr Prot Food Analyt Chem. 1:F4.2.1-F4.2.6.
Lin W, Shuai B, Springer P (2003) The Arabidopsis LATERAL ORGAN BOUNDARIES–domain gene ASYMMETRIC LEAVES2 functions in the repression of KNOX gene expression and in adaxial abaxial patterning. Plant Cell. 15:2241–2252.
Lopez-Galvez F, Ragaert P, Palermo LA, Eriksson M, Devlieghere F (2013) Effect of new sanitizing formulations on quality of fresh-cut iceberg lettuce. Postharvest Biol Tec. 85: 102-108.
Muhammad T, Zhang J, Ma Y et al (2019) Overexpression of a mitogen-activated protein kinase SlMAPK3 positively regulates tomato tolerance to cadmium and drought stress. Molecules. 24:556.
Norberg M, Holmlund M, Nilsson O (2005). The BLADE ON PETIOLE genes act redundantly to control the growth and development of lateral organs. Development. 132: 2203–2213.
Ori N, Eshed Y, Chuck G, Bowman JL, Hake S (2000) Mechanisms that control knox gene expression in the Arabidopsis shoot. Development. 127: 5523–5532.
Patel A, Patel A, Patel A, Patel NM (2010) Determination of polyphenols and free radical scavenging activity of Tephrosia purpurea linn leaves (Leguminosae). Pharmacogn Res. 2: 152–158.
Pieterse CM, Van Loon LC (2004) NPR1: The spider in the web of induced resistance signaling pathways. Curr Opin Plant Biol. 7:456–464.
Schelbert S, Aubry S, Burla B, Agne B, Kessler F, Krupinska K, Ho S (2009) Pheophytin Pheophorbide hydrolase (Pheophytinase) is Involved in chlorophyll breakdown during leaf senescence in Arabidopsis. Plant Cell. 21:767–785.
Krishna R, Karkute SG, Ansari WA, Jaiswal DK, Verma JP, Singh M (2019) Transgenic tomatoes for abiotic stress tolerance: status and way ahead. 3 Biotech. 9: 143.
Sedgwick SG, Smerdon SJ (1999) The ankyrin repeat: A diversity of interactions on a common structural framework. Trends Biochem Sci. 24:311–316.
Semiarti E, Ueno Y, Tsukaya H, Iwakaka H, Machida C, Machida Y (2001) The ASYMMETRIC LEAVES2 gene of Arabidopsis thaliana regulates formation of a symmetric lamina, establishment of venation and repression of meristem-related homeobox genes in leaves. Development. 128:1771–1783.
Singh Z, Khan AS, Zhu S, Payne AD (2013) Nitric oxide in the regulation of fruit ripening: challenges and thrusts. Stewart Postharvest Rev. 9:1-11.
Sukharev SI, Klenchin VA, Serov SM, Chernomordik LV, Chizmadzhev YuA (1992) Electroporation and electrophoretic DNA transfer into cells. The effect of DNA interaction with electropores. Biophys J. 63:1320-1328.
Sun Y, Zhou Q, Zhang W, Fu Y, Huang H (2002) ASYMMETRIC LEAVES1, an Arabidopsis gene that is involved in the control of cell differentiation in leaves. Planta. 214:694–702.
Thi Man Le, Thi Lan Huong Phung and Phi Bang Cao (2020) Effect of magnesium on growth, fruit yield and some biochemical indices of hydroponic black tomato. Asian J Plant Sci. 19: 273-278.
Uchida N, Kimura S, Koenig D, Sinha N (2010) Coordination of leaf development via regulation of KNOX1 genes. J Plant Res. 123:7-17.
Wang WR, Liang JH, Wang GF, Sun MX, Peng FT, Xiao YS (2020) Overexpression of PpSnRK1 in tomato enhanced salt tolerance by regulating ABA signaling pathway and reactive oxygen metabolism. BMC Plant Biol. 20:128.
Zhang Y, Butelli E, De Stefano R, Schoonbeek HJ, Magusin A, Pagliarani C, Wellner N, Hill L, Orzaez D, Granell A, Jones JD, Martin C (2013) Anthocyanins double the shelf life of tomatoes by delaying overripening and reducing susceptibility to gray mold. Curr Biol. 23:1094–1100.
Zhang N, Roberts HM, Van Eck J, Martin GB (2020) Generation and Molecular Characterization of CRISPR/Cas9-Induced Mutations in 63 Immunity-Associated Genes in Tomato Reveals Specificity and a Range of Gene Modifications. Front Plant Sci. 11:10.